data |> glimpse()
#> Rows: 39
#> Columns: 92
#> $ id <chr> "9e858cd8", "a053ce64", "db850249", "5af4…
#> $ timestamp <dttm> 2023-05-21 11:18:49, 2023-05-26 17:55:52…
#> $ birth_date <date> 1991-03-22, 2001-07-25, 1998-01-12, 2001…
#> $ age <dbl> 32.16388889, 21.83611111, 25.47222222, 22…
#> $ weight_before <dbl> 70, 50, 70, 84, 49, 60, 66, 73, 52, 68, 6…
#> $ weight_current <dbl> 86, 75, 76, 87, 58, 91, 79, 90, 62, 80, 7…
#> $ height <dbl> 160, 156, 169, 168, 157, 164, 159, 159, 1…
#> $ bmi_before <dbl> 27.34375000, 20.54569362, 24.50894577, 29…
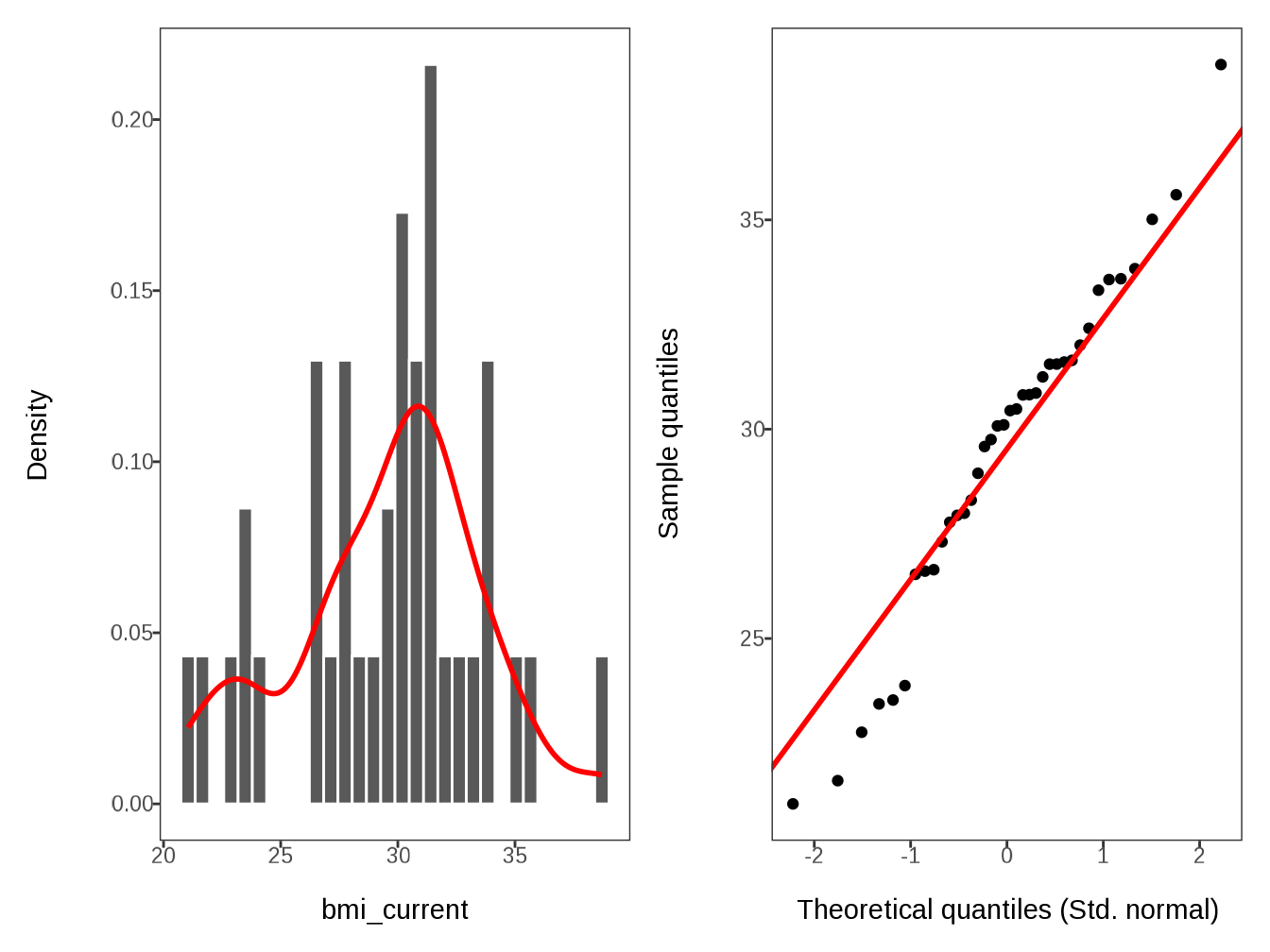
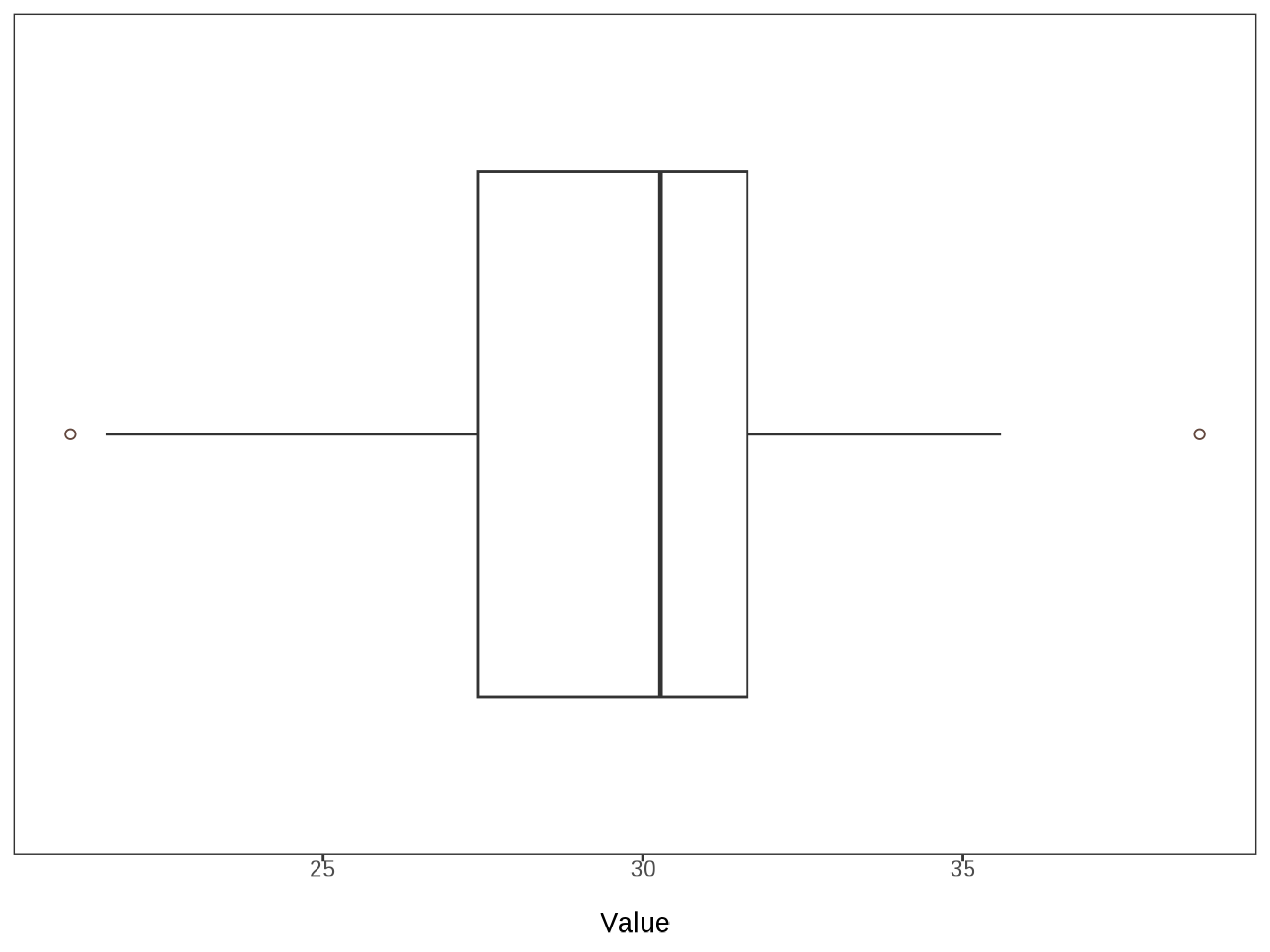
#> $ bmi_current <dbl> 33.59375000, 30.81854043, 26.60971255, 30…
#> $ family_income <dbl> 4000, 2800, 5000, 4000, 3000, 9000, 2500,…
#> $ gestations <dbl> 1, 1, 1, 1, 1, 1, 1, 2, 1, 2, 1, 4, 1, 1,…
#> $ study <lgl> FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, …
#> $ work <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, FALSE, FALS…
#> $ health_plan <lgl> FALSE, FALSE, TRUE, FALSE, FALSE, FALSE, …
#> $ solo <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,…
#> $ ethnicity <ord> Branca, Parda, Branca, Parda, Branca, Par…
#> $ exercise <lgl> NA, NA, NA, NA, NA, NA, NA, FALSE, FALSE,…
#> $ education <ord> NA, NA, NA, NA, NA, NA, NA, Ensino superi…
#> $ deliveries <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0,…
#> $ abortions <dbl> 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 2, 0, 0,…
#> $ birth_center_delivery <lgl> FALSE, TRUE, TRUE, FALSE, FALSE, FALSE, T…
#> $ ga_start <date> 2022-09-03, 2022-09-03, 2022-10-13, 2022…
#> $ partograph_begin <dttm> NA, 2023-06-08 11:40:00, 2023-07-11 15:0…
#> $ begin_dilation <int> NA, 7, 5, NA, NA, NA, 7, 7, 7, NA, 10, NA…
#> $ baby_birth <dttm> NA, 2023-06-08 14:54:00, 2023-07-11 18:0…
#> $ baby_weight <dbl> NA, 2995, 3010, NA, NA, NA, 3730, 2740, 3…
#> $ baby_length <dbl> NA, 46.0, 45.5, NA, NA, NA, 51.0, 47.0, 4…
#> $ baby_head_perimeter <dbl> NA, 32.5, 33.0, NA, NA, NA, 34.0, 32.0, 3…
#> $ baby_thoracic_perimeter <dbl> NA, 31.0, 32.0, NA, NA, NA, 34.5, 31.0, 3…
#> $ baby_abdominal_perimeter <dbl> NA, 30.0, 33.5, NA, NA, NA, 31.5, 28.0, 3…
#> $ apgar_score_1_minute <int> NA, 9, 9, NA, NA, NA, 9, 9, 9, NA, 9, NA,…
#> $ apgar_score_5_minute <int> NA, 10, 10, NA, NA, NA, 10, 10, 10, NA, 1…
#> $ partograph_duration <Duration> NA, 11640s (~3.23 hours), 10920s (~3…
#> $ n_primary_sleep <int> 25, 13, 7, 23, 18, 16, 22, 13, 17, 20, 17…
#> $ his_mean <time> 23:48:01.600000, 02:24:28.846154, 01:28:…
#> $ his_sd <time> 01:06:59.308896, 01:26:16.088514, 01:06:…
#> $ his_min <time> 21:58:04, 00:04:15, 00:05:52, 22:47:31, …
#> $ his_max <time> 26:42:04, 05:58:15, 03:41:52, 28:21:31, …
#> $ hfs_mean <time> 06:55:47.200000, 10:53:47.307692, 09:30:…
#> $ hfs_sd <time> 01:45:15.303318, 02:30:46.599871, 00:59:…
#> $ hfs_min <time> 03:27:04, 07:38:15, 08:02:52, 06:57:31, …
#> $ hfs_max <time> 10:32:04, 15:37:15, 10:24:52, 13:54:31, …
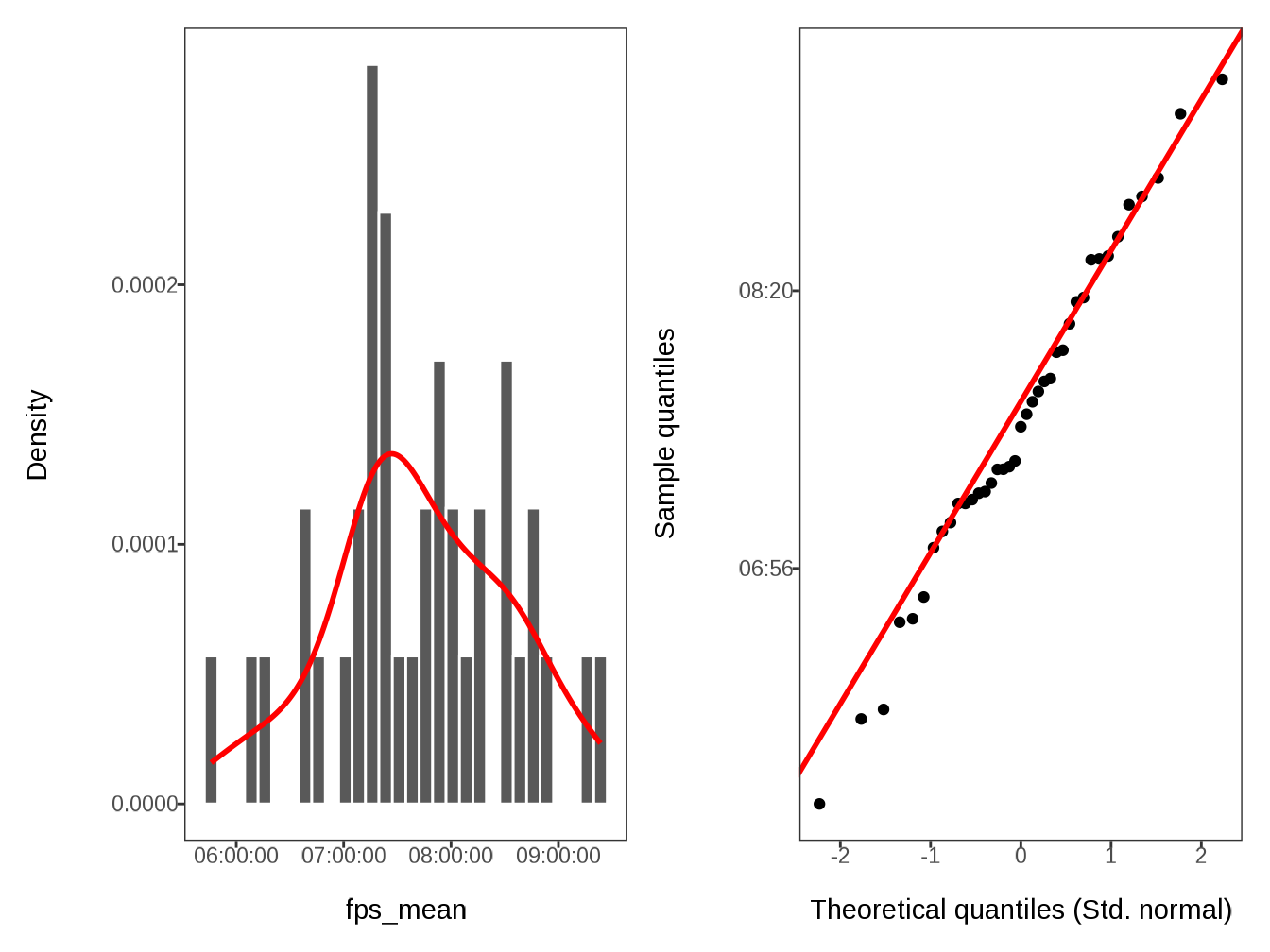
#> $ fps_mean <time> 07:07:45.600000, 08:29:18.461538, 08:01:…
#> $ fps_sd <time> 02:00:10.727009, 02:48:00.011447, 01:15:…
#> $ fps_min <time> 03:07:00, 04:22:00, 06:34:00, 06:14:00, …
#> $ fps_max <time> 10:41:00, 13:40:00, 09:54:00, 12:14:00, …
#> $ tts_mean <time> 05:26:28.800000, 06:54:23.076923, 06:50:…
#> $ tts_sd <time> 01:29:47.934298, 02:08:15.357242, 01:07:…
#> $ tts_min <time> 02:47:00, 03:59:00, 05:47:00, 05:37:00, …
#> $ tts_max <time> 08:07:00, 10:00:00, 08:35:00, 09:34:00, …
#> $ waso_mean <time> 01:41:16.800000, 01:34:55.384615, 01:11:…
#> $ waso_sd <time> 01:01:08.086149, 00:56:56.617761, 00:41:…
#> $ waso_min <time> 00:14:00, 00:23:00, 00:28:00, 00:28:00, …
#> $ waso_max <time> 04:04:00, 03:40:00, 02:34:00, 04:03:00, …
#> $ tts_fps_mean <dbl> 0.7758617755, 0.8202364594, 0.8544758095,…
#> $ tts_fps_sd <dbl> 0.11565879087, 0.06542581849, 0.076444496…
#> $ tts_fps_min <dbl> 0.5000000000, 0.7317073171, 0.6986301370,…
#> $ tts_fps_max <dbl> 0.9482758621, 0.9238095238, 0.9289340102,…
#> $ awakenings_mean <dbl> 8.440000000, 17.230769231, 10.285714286, …
#> $ awakenings_sd <dbl> 3.937850513, 7.350213672, 3.903600292, 3.…
#> $ awakenings_min <dbl> 2, 6, 3, 8, 1, 10, 10, 4, 7, 0, 4, 8, 4, …
#> $ awakenings_max <dbl> 17, 34, 15, 23, 14, 29, 27, 20, 23, 7, 14…
#> $ n_secondary_sleep <int> 35, 10, 5, 1, 8, 9, 18, 5, 2, 12, 19, NA,…
#> $ his_mean_secondary <time> 24:56:58.857143, 15:33:03.000000, 20:21:…
#> $ his_sd_secondary <time> 08:57:33.700323, 02:37:02.687515, 08:22:…
#> $ his_min_secondary <time> 12:01:04, 12:00:15, 13:22:52, 14:15:31, …
#> $ his_max_secondary <time> 35:07:04, 21:21:15, 34:11:52, 14:15:31, …
#> $ hfs_mean_secondary <time> 35:11:04.000000, 35:48:15.000000, 35:54:…
#> $ hfs_sd_secondary <time> 05:37:54.966309, 08:01:22.049789, 07:46:…
#> $ hfs_min_secondary <time> 18:10:04, 19:15:15, 22:29:52, 15:43:31, …
#> $ hfs_max_secondary <time> 41:45:04, 41:45:15, 41:11:52, 15:43:31, …
#> $ fps_mean_secondary <time> 01:19:13.714286, 01:03:12.000000, 01:09:…
#> $ fps_sd_secondary <time> 01:00:34.950510, 00:55:11.048172, 01:12:…
#> $ fps_min_secondary <time> 00:14:00, 00:09:00, 00:11:00, 01:28:00, …
#> $ fps_max_secondary <time> 03:35:00, 02:57:00, 03:03:00, 01:28:00, …
#> $ tts_mean_secondary <time> 01:14:56.571429, 00:55:42.000000, 01:04:…
#> $ tts_sd_secondary <time> 00:57:47.501171, 00:45:55.220499, 01:04:…
#> $ tts_min_secondary <time> 00:14:00, 00:09:00, 00:11:00, 01:16:00, …
#> $ tts_max_secondary <time> 03:32:00, 02:24:00, 02:41:00, 01:16:00, …
#> $ waso_mean_secondary <time> 00:04:17.142857, 00:07:30.000000, 00:05:…
#> $ waso_sd_secondary <time> 00:05:45.462788, 00:10:29.444199, 00:09:…
#> $ waso_min_secondary <time> 00:00:00, 00:00:00, 00:00:00, 00:12:00, …
#> $ waso_max_secondary <time> 00:25:00, 00:33:00, 00:22:00, 00:12:00, …
#> $ tts_fps_mean_secondary <dbl> 0.9555844698, 0.9118161628, 0.9546406611,…
#> $ tts_fps_sd_secondary <dbl> 0.05273858710, 0.07690741832, 0.058366343…
#> $ tts_fps_min_secondary <dbl> 0.8000000000, 0.7916666667, 0.8797814208,…
#> $ tts_fps_max_secondary <dbl> 1.0000000000, 1.0000000000, 1.0000000000,…
#> $ awakenings_mean_secondary <dbl> 0.9714285714, 1.6000000000, 1.2000000000,…
#> $ awakenings_sd_secondary <dbl> 1.0706159383, 1.6465452047, 1.6431676725,…
#> $ awakenings_min_secondary <dbl> 0, 0, 0, 2, 0, 0, 0, 0, 1, 0, 0, NA, NA, …
#> $ awakenings_max_secondary <dbl> 4, 5, 4, 2, 3, 7, 4, 5, 1, 1, 5, NA, NA, …
#> $ ga_duration <Duration> NA, 24083640s (~39.82 weeks), 234901…