data |> glimpse()
#> Rows: 8
#> Columns: 62
#> $ id <chr> "a053ce64", "b0e857dd", "1d215e39", "b37bd…
#> $ birth_center_delivery <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, …
#> $ labor_start <dttm> 2023-08-06 09:30:00, 2023-08-01 03:30:00,…
#> $ labor_duration <Duration> 19440s (~5.4 hours), 99300s (~1.15 da…
#> $ timestamp <dttm> 2023-05-26 17:55:52, 2023-07-16 22:02:13,…
#> $ birth_date <date> 2001-07-25, 1989-03-06, 1996-05-11, 1989-…
#> $ age <dbl> 21.83611111, 34.36111111, 27.20277778, 34.…
#> $ weight_before <dbl> 50, 75, 66, 87, 70, 73, 52, 61
#> $ weight_current <dbl> 75, 89, 79, 94, 76, 90, 62, 77
#> $ height <dbl> 156, 164, 159, 174, 169, 159, 148, 160
#> $ bmi_before <dbl> 20.54569362, 27.88518739, 26.10656224, 28.…
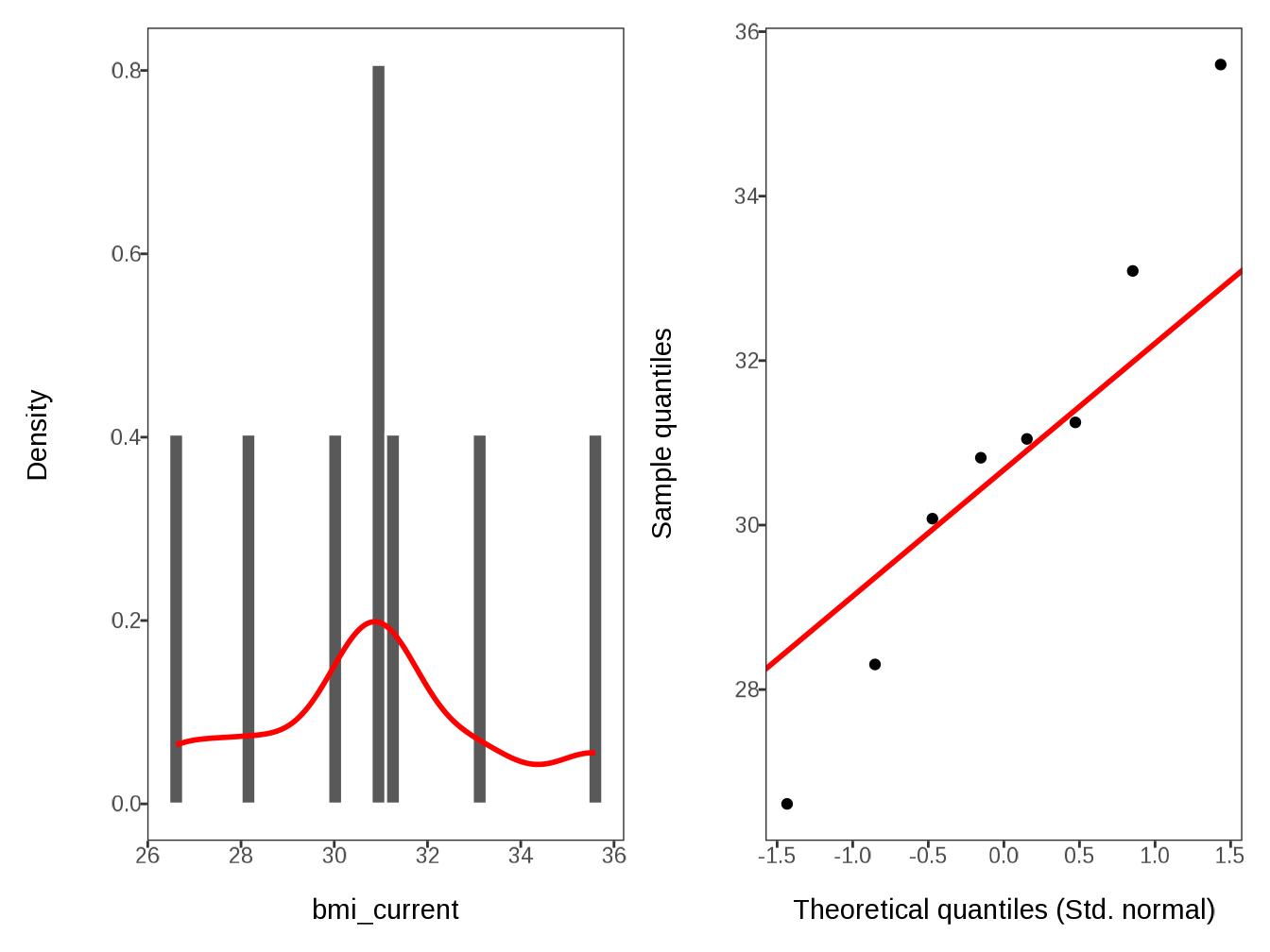
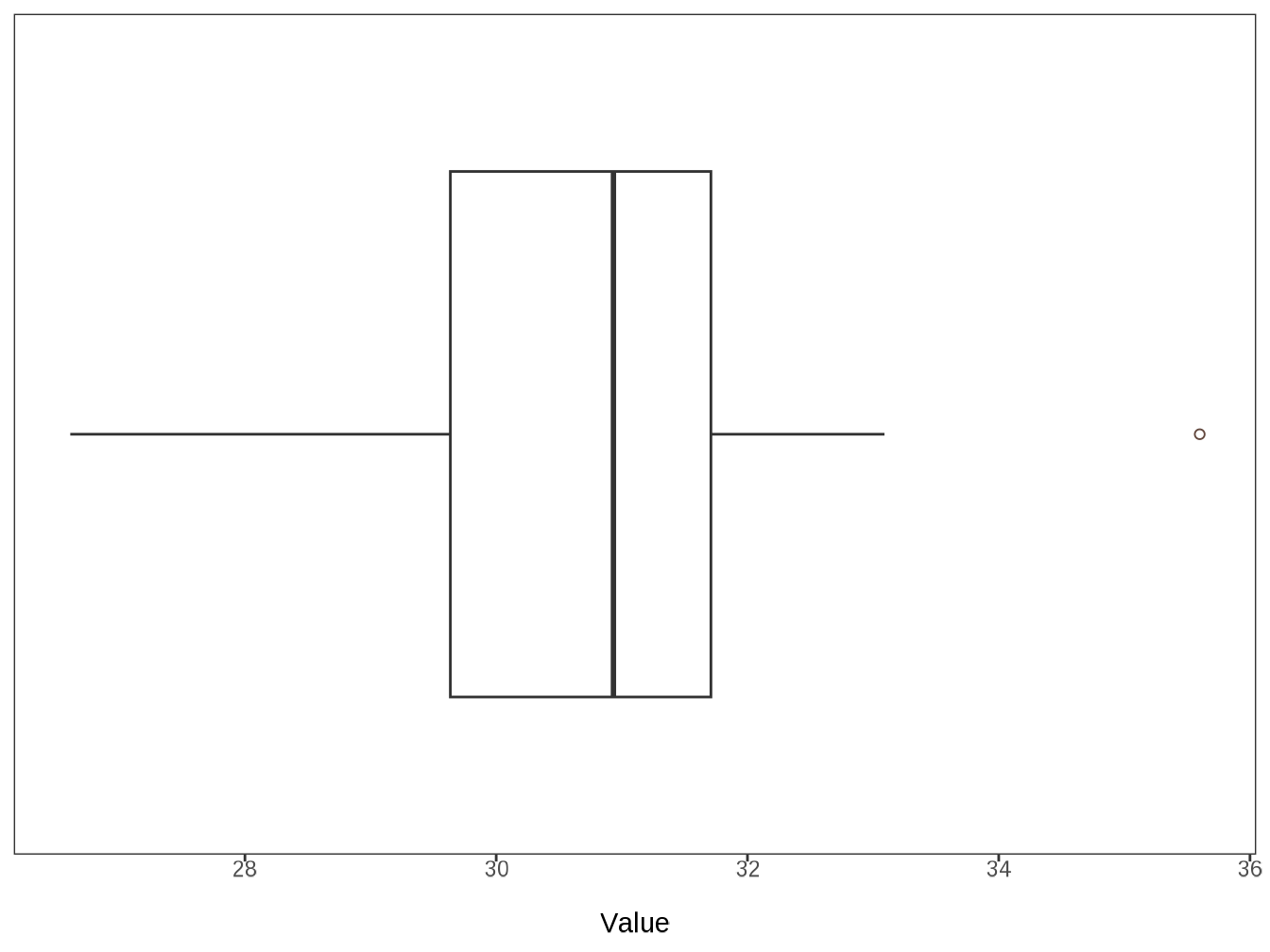
#> $ bmi_current <dbl> 30.81854043, 33.09042237, 31.24876389, 31.…
#> $ family_income <dbl> 2800, 6000, 2500, 6000, 5000, 3000, 3500, …
#> $ gestations <dbl> 1, 1, 1, 1, 1, 2, 1, 1
#> $ study <lgl> TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, F…
#> $ work <lgl> TRUE, TRUE, FALSE, TRUE, TRUE, FALSE, TRUE…
#> $ health_plan <lgl> FALSE, TRUE, FALSE, TRUE, TRUE, FALSE, FAL…
#> $ solo <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, …
#> $ ethnicity <ord> Parda, Branca, Parda, Branca, Branca, Bran…
#> $ exercise <lgl> NA, NA, NA, NA, NA, FALSE, FALSE, FALSE
#> $ education <ord> NA, NA, NA, NA, NA, Ensino superior incomp…
#> $ deliveries <dbl> 0, 0, 0, 0, 0, 0, 0, 0
#> $ abortions <dbl> 0, 0, 0, 0, 0, 1, 0, 0
#> $ his_min <time> 01:26:15, 22:30:44, 23:25:39, 00:01:18, 00…
#> $ his_max <time> 03:41:15, 05:55:44, 01:38:39, 02:05:18, 03…
#> $ his_mean <time> 02:18:45, 00:39:32, 00:37:19, 01:12:54, 01…
#> $ his_sd <time> 00:37:56, 02:40:43, 00:51:14, 00:38:38, 01…
#> $ hfs_min <time> 07:42:15, 04:02:44, 07:13:39, 05:55:18, 08…
#> $ hfs_max <time> 15:37:15, 10:32:44, 10:51:39, 09:13:18, 10…
#> $ hfs_mean <time> 11:10:40, 07:30:20, 08:36:59, 07:51:18, 09…
#> $ hfs_sd <time> 02:25:50, 02:13:33, 01:13:28, 01:10:06, 0…
#> $ fps_min <time> 05:01:00, 02:38:00, 06:21:00, 05:33:00, 0…
#> $ fps_max <time> 13:40:00, 10:57:00, 09:44:00, 07:15:00, 0…
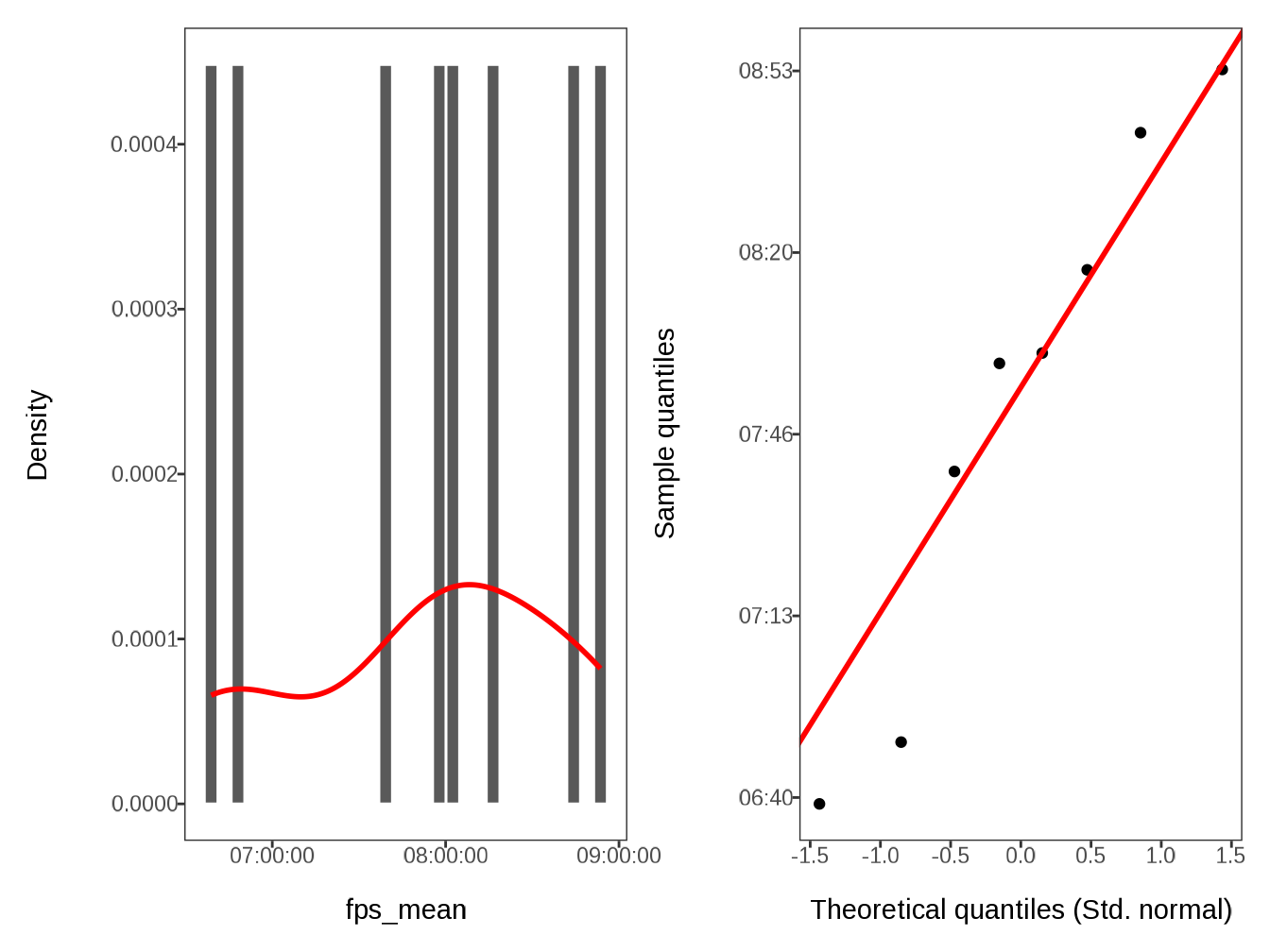
#> $ fps_mean <time> 08:42:00, 06:50:10, 07:59:40, 06:38:50, 0…
#> $ fps_sd <time> 02:49:52, 03:10:58, 01:15:16, 00:39:54, 0…
#> $ tts_min <time> 04:05:00, 02:20:00, 03:57:00, 05:11:00, 0…
#> $ tts_max <time> 10:00:00, 08:31:00, 06:23:00, 06:44:00, 0…
#> $ tts_mean <time> 06:56:37, 05:47:36, 05:19:00, 06:00:00, 0…
#> $ tts_sd <time> 02:05:55, 02:29:07, 00:49:14, 00:33:49, 0…
#> $ tts_fps_min <dbl> 73.17, 77.78, 53.74, 84.60, 69.86, 73.33,…
#> $ tts_fps_max <dbl> 92.38, 93.43, 80.35, 95.96, 92.89, 95.11,…
#> $ tts_fps_mean <dbl> 80.78, 86.06, 67.11, 90.53, 85.45, 85.62,…
#> $ tts_fps_sd <dbl> 6.34, 5.14, 9.20, 3.68, 7.08, 8.32, 3.49,…
#> $ waso_min <time> 00:40:00, 00:18:00, 01:29:00, 00:17:07, 0…
#> $ waso_max <time> 03:40:00, 02:26:00, 04:08:00, 01:07:00, 0…
#> $ waso_mean <time> 01:45:25, 01:03:12, 02:40:00, 00:38:20, 0…
#> $ waso_sd <time> 01:00:18, 00:48:39, 00:58:46, 00:17:07, 00…
#> $ awakenings_min <dbl> 8, 4, 10, 3, 3, 8, 8, 6
#> $ awakenings_max <dbl> 34, 13, 21, 10, 15, 20, 19, 14
#> $ awakenings_mean <dbl> 18.00, 8.40, 16.67, 5.83, 10.29, 13.80, 13…
#> $ awakenings_sd <dbl> 7.33, 3.50, 4.27, 2.27, 3.61, 4.26, 4.31, …
#> $ partograph_begin <dttm> 2023-06-08 11:40:00, 2023-08-02 03:00:00, …
#> $ begin_dilation <dbl> 7, 7, 7, 6, 5, 7, 7, 10
#> $ baby_birth <dttm> 2023-06-08 14:54:00, 2023-08-02 07:05:00,…
#> $ baby_weight <dbl> 2995, 2785, 3730, 3340, 3010, 2740, 3080,…
#> $ baby_length <dbl> 46.0, 46.0, 51.0, 49.5, 45.5, 47.0, 49.0,…
#> $ baby_head_perimeter <dbl> 32.5, 33.0, 34.0, 35.0, 33.0, 32.0, 31.2,…
#> $ baby_thoracic_perimeter <dbl> 31.0, 32.0, 34.5, 33.5, 32.0, 31.0, 32.0, …
#> $ baby_abdominal_perimeter <dbl> 30.0, 30.0, 31.5, 32.5, 33.5, 28.0, 30.0, …
#> $ apgar_score_1_minute <int> 9, 9, 9, 9, 9, 9, 9, 9
#> $ apgar_score_5_minute <int> 10, 10, 10, 10, 10, 10, 10, 10
#> $ waso_prop <dbl> 0.20194763729, 0.15408370581, 0.3335649756…